
ggExametrika provides ggplot2-based visualization for the exametrika package. It supports a wide range of psychometric models:
| Model | Description |
|---|---|
| IRT | Item Response Theory (2PL, 3PL, 4PL) |
| GRM | Graded Response Model |
| LCA | Latent Class Analysis |
| LRA | Latent Rank Analysis |
| LRAordinal | Latent Rank Analysis for ordinal data |
| LRArated | Latent Rank Analysis for rated data |
| Biclustering | Simultaneous item/student clustering (binary) |
| nominalBiclustering | Biclustering for nominal data |
| ordinalBiclustering | Biclustering for ordinal data |
| IRM | Infinite Relational Model |
| LDLRA | Locally Dependent Latent Rank Analysis |
| LDB | Locally Dependent Biclustering |
| BINET | Bayesian Network and Test |
| BNM | Bayesian Network Model |
Shojima, Kojiro (2022) Test Data Engineering: Latent Rank Analysis, Biclustering, and Bayesian Network (Behaviormetrics: Quantitative Approaches to Human Behavior, 13), Springer, ISBN 978-981-16-9985-6
# install.packages("devtools")
devtools::install_github("kosugitti/ggExametrika")All plot functions take exametrika output directly and return ggplot
objects. Functions are named plotXXX_gg().
library(exametrika)
library(ggExametrika)
result_irt <- IRT(J15S500, model = 3)
plots <- plotICC_gg(result_irt)
plots[[5]]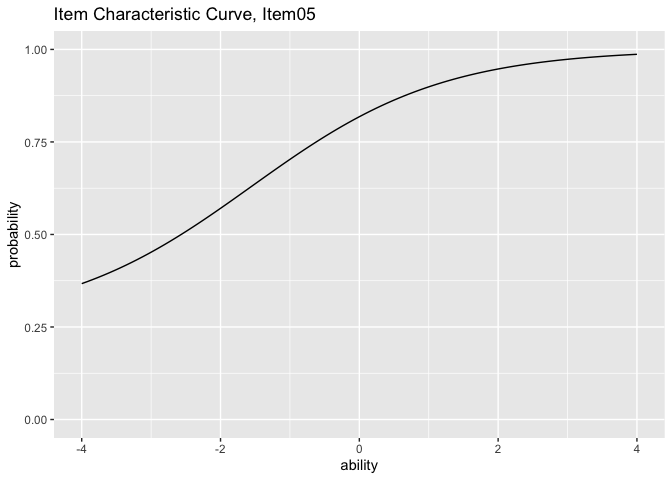
combinePlots_gg(plots)
# All ICCs on a single plot
plotICC_overlay_gg(result_irt, show_legend = TRUE)
# All IICs on a single plot (also works with GRM)
plotIIC_overlay_gg(result_irt, items = c(1, 3, 5), show_legend = TRUE)plots <- plotIIC_gg(result_irt)
combinePlots_gg(plots, selectPlots = 8:11)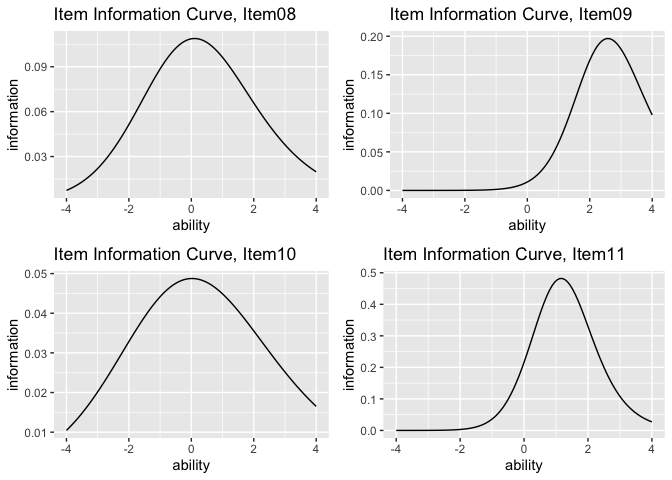
plotTIC_gg(result_irt)
plotTRF_gg(result_irt)result_grm <- GRM(J5S1000)
plots <- plotICRF_gg(result_grm)
plots[[1]]
combinePlots_gg(plots, selectPlots = 1:5)
# GRM also supports IIC and TIC
plotIIC_gg(result_grm)
plotTIC_gg(result_grm)result_lca <- LCA(J15S500, ncls = 3)
plotIRP_gg(result_lca) # Item Reference Profile
plotFRP_gg(result_lca) # Field Reference Profile
plotTRP_gg(result_lca) # Test Reference Profile
plotLCD_gg(result_lca) # Latent Class Distribution
plotCMP_gg(result_lca) # Class Membership Profileresult_lra <- LRA(J15S500, nrank = 4)
plotIRP_gg(result_lra) # Item Reference Profile
plotFRP_gg(result_lra) # Field Reference Profile
plotTRP_gg(result_lra) # Test Reference Profile
plotLRD_gg(result_lra) # Latent Rank Distribution
plotRMP_gg(result_lra) # Rank Membership Profileresult_lra_ord <- LRA(J5S1000, nrank = 4) # ordinal data
plotScoreFreq_gg(result_lra_ord) # Score Frequency Distribution
plotScoreRank_gg(result_lra_ord) # Score-Rank Heatmap
plotICRP_gg(result_lra_ord) # Item Category Reference Profile
plotICBR_gg(result_lra_ord) # Item Category Boundary Response (ordinal only)
plotRMP_gg(result_lra_ord) # Rank Membership Profileresult_bic <- Biclustering(J35S515, nfld = 5, nrank = 6)
plotFRP_gg(result_bic) # Field Reference Profile
plotTRP_gg(result_bic) # Test Reference Profile
plotLCD_gg(result_bic) # Latent Class Distribution
plotLRD_gg(result_bic) # Latent Rank Distribution
plotCMP_gg(result_bic) # Class Membership Profile
plotRMP_gg(result_bic) # Rank Membership Profile
plotCRV_gg(result_bic) # Class Reference Vector
plotRRV_gg(result_bic) # Rank Reference Vector
plotArray_gg(result_bic) # Array Plot (heatmap)# Nominal Biclustering
result_nom <- Biclustering(data, ncls = 3, nfld = 4)
plotFRP_gg(result_nom, stat = "mean") # stat: "mean", "median", or "mode"
plotFCRP_gg(result_nom, style = "line") # Field Category Response Profile (style: "line" or "bar")
plotScoreField_gg(result_nom) # Expected Score Heatmap (field x class/rank)
plotCRV_gg(result_nom, stat = "mean") # Class Reference Vector
plotRRV_gg(result_nom, stat = "mean") # Rank Reference Vector
plotArray_gg(result_nom) # Array Plot
# Ordinal Biclustering (additional)
plotFCBR_gg(result_ord) # Field Cumulative Boundary Reference (ordinal only)result_ldb <- LDB(J35S515, ncls = 6, nfld = 5)
plotFRP_gg(result_ldb) # Field Reference Profile
plotTRP_gg(result_ldb) # Test Reference Profile
plotLRD_gg(result_ldb) # Latent Rank Distribution
plotRMP_gg(result_ldb) # Rank Membership Profile
plotArray_gg(result_ldb) # Array Plot
plotFieldPIRP_gg(result_ldb) # Field Parent Item Reference Profile
plotGraph_gg(result_ldb) # DAG per rankresult_binet <- BINET(J35S515, ncls = 6, nfld = 5)
plotFRP_gg(result_binet) # Field Reference Profile
plotTRP_gg(result_binet) # Test Reference Profile
plotLRD_gg(result_binet) # Latent Rank Distribution
plotRMP_gg(result_binet) # Rank Membership Profile
plotArray_gg(result_binet) # Array Plot
plotGraph_gg(result_binet, show_edge_label = TRUE) # DAG with edge labelsresult_bnm <- BNM(J15S500)
plotGraph_gg(result_bnm)
result_ldlra <- LDLRA(J15S500, ncls = 5)
plotGraph_gg(result_ldlra) # One DAG per rank| Function | IRT | GRM |
|---|---|---|
| plotICC_gg | x | |
| plotICC_overlay_gg | x | |
| plotIIC_gg | x | x |
| plotIIC_overlay_gg | x | x |
| plotTIC_gg | x | x |
| plotTRF_gg | x | |
| plotICRF_gg | x |
| Function | LCA | LRA | LRAordinal | LRArated |
|---|---|---|---|---|
| plotIRP_gg | x | x | ||
| plotFRP_gg | x | x | ||
| plotTRP_gg | x | x | ||
| plotLCD_gg | x | |||
| plotLRD_gg | x | |||
| plotCMP_gg | x | |||
| plotRMP_gg | x | x | x | |
| plotScoreFreq_gg | x | x | ||
| plotScoreRank_gg | x | x | ||
| plotICRP_gg | x | x | ||
| plotICBR_gg | x |
| Function | Bic. | nomBic. | ordBic. | IRM |
|---|---|---|---|---|
| plotFRP_gg | x | x | x | x |
| plotTRP_gg | x | x | ||
| plotLCD_gg | x | x | x | |
| plotLRD_gg | x | x | x | |
| plotCMP_gg | x | x | x | |
| plotRMP_gg | x | x | ||
| plotCRV_gg | x | x | x | |
| plotRRV_gg | x | x | x | |
| plotArray_gg | x | x | x | x |
| plotFCRP_gg | x | x | ||
| plotFCBR_gg | x | |||
| plotScoreField_gg | x | x |
| Function | LDLRA | LDB | BINET | BNM |
|---|---|---|---|---|
| plotIRP_gg | x | |||
| plotFRP_gg | x | x | ||
| plotTRP_gg | x | x | ||
| plotLRD_gg | x | x | x | |
| plotRMP_gg | x | x | x | |
| plotArray_gg | x | x | ||
| plotFieldPIRP_gg | x | |||
| plotGraph_gg | x | x | x | x |
| Function | Description |
|---|---|
| combinePlots_gg | Arrange multiple plots in a grid |
All plot functions support these customization options:
| Parameter | Description | Default |
|---|---|---|
title |
TRUE (auto), FALSE (none), or character
string |
TRUE |
colors |
Color vector (colorblind-friendly default) | auto |
linetype |
"solid", "dashed", "dotted",
etc. |
"solid" |
show_legend |
Show/hide legend | TRUE |
legend_position |
"right", "top", "bottom",
"left" |
"right" |
Some functions have additional parameters:
| Parameter | Functions | Description |
|---|---|---|
stat |
plotFRP_gg, plotCRV_gg, plotRRV_gg | "mean", "median", or "mode"
for polytomous data |
style |
plotFCRP_gg | "line" or "bar" |
show_labels |
plotRRV_gg | Show value labels (uses ggrepel) |